Nobel Prize Day 2023: The Road to mRNA Vaccines
December 10th is Nobel Prize Day, and this year the Prize for Physiology or Medicine went to two scientists whose work directly impacted all of us these last couple years: Drs. Katalin Karikó and Drew Weissman. Karikó and Weissman’s research into how mRNA interacts with our immune systems enabled the astonishingly rapid development of vaccines against the SARS-CoV-2 virus.

A very brief history of vaccines
Scientists have been systematically studying the body’s natural immune response to pathogens in order to develop vaccines since the late 1700s, when Edward Jenner and others successfully created a vaccine for the smallpox virus. However, the general practice of introducing small amounts of pathogens to healthy people in order to confer immunity may go back even farther.
After Jenner, the first century of vaccine usage employed small amounts of whole viruses – living or dead – to stimulate the production of antibodies against that specific pathogen. More recently, vaccines have used just a piece of the virus, such as surface proteins or part of its genetic code embedded in a harmless viral vector, to more safely achieve the same aim. All of these methods of vaccine production are relatively slow and resource-intensive, as they are produced through large-scale cell culture.

Where does mRNA come in?
In an organism, messenger RNA (mRNA) acts as a transient intermediary between DNA and protein. Information encoded in DNA is transcribed into mRNA, which serves as instructions for assembling proteins. By the late 20th century, mRNA could be produced with relative ease and high throughput via a process called in vitro transcription – or transcription in a dish. In theory, in vitro-transcribed mRNA could be used to assemble the viral proteins required for some forms of vaccine in a more efficient way compared to cell culture. However, this posed new problems: mRNA is not particularly stable outside of a cell, nor is it welcomed by an organism’s immune system. The need for a sophisticated mRNA delivery system, along with the adverse reactions seen in animal models, limited its applications in medicine.
How did Drs. Karikó and Weissman change the use of mRNA in medicine?
In their work at the University of Pennsylvania, Karikó and Weissman investigated the interactions between mRNA and dendrites – immune cells that are key modulators in how an organism responds to pathogens, and consequently vaccines. They found that in vitro-transcribed mRNA was recognized as foreign pathogenic material by dendrites, while cell-derived mRNA was not. To understand why, they looked at the underlying molecular composition of both forms of mRNA. In mammalian cell transcription, mRNA often undergoes certain chemical alterations that do not occur during in vitro transcription. In 2005, Karikó and Weissman published a study showing that chemically modified in vitro-derived mRNA did not produce the adverse immune response triggered by unmodified mRNA, overcoming a major barrier to therapeutic applications.
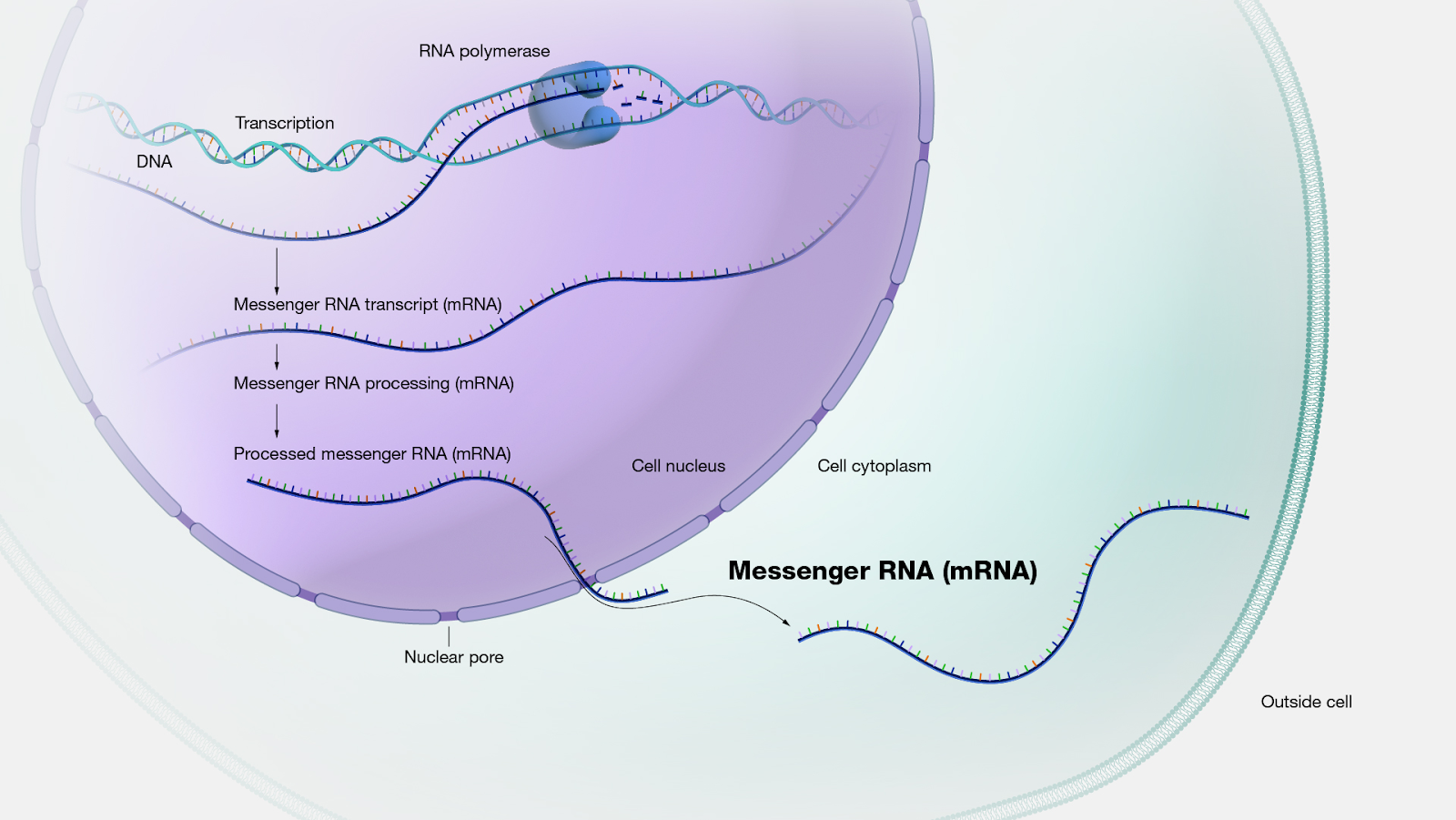
How mRNA is produced in a eukaryotic cell. Figure from the National Human Genome Research Institute.
In mammalian cells, mRNA is made up of four bases: adenine (A), guanine (G), cytosine (C), and uracil (U). By modifying uridine into a form called pseudouridine, the researchers went on to make other critical discoveries in 2008 and 2010, respectively: modified in vitro-transcribed mRNA has increased translational capacity, and is not as readily degraded by the protein kinase R enzyme (PKR). All of these findings made modified mRNA a promising candidate as a vaccine vector:
- It could be used to robustly trigger the production of viral proteins
- It was biologically and chemically stable
- It would not induce an adverse inflammatory response
- It could be rapidly produced at-scale without cell culture
The last point was key in allowing scientists and regulators to develop safe and effective vaccines at the record pace seen during the pandemic.
The future of vaccine development
Not only can mRNA vaccines be developed and deployed much more quickly than their predecessors, but they’re also fundamentally more versatile. Instead of needing an existing virus or pieces of it to develop a vaccine, mRNA technology employs genetic code to tell the patient’s body which proteins to produce. In order to create a vaccine for a new pathogen, scientists could simply change the code. Researchers are already working on developing mRNA vaccines for other infectious diseases including HIV, malaria, and tuberculosis.
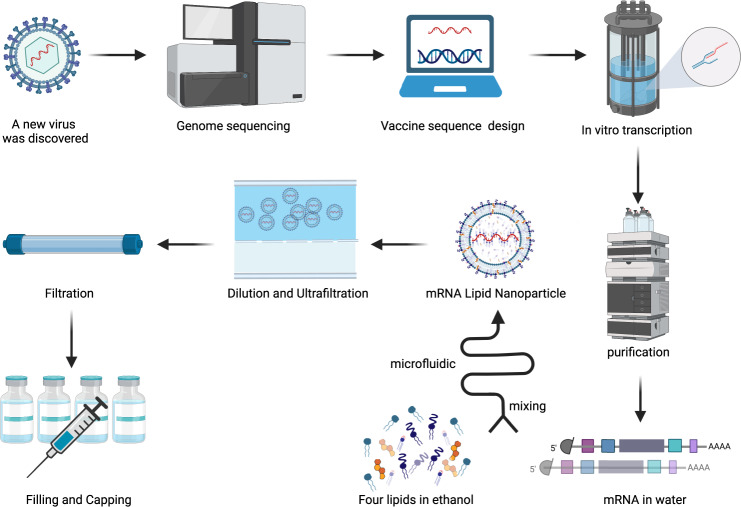
Furthermore, mRNA vaccination is being explored as a route to prevent and treat a host of other non-infectious diseases, such as allergies, genetic disorders, and even cancer. Integrating large-scale datasets like gnomAD will be critical in ensuring the safety and efficacy of these approaches across different patient populations.

